Project A8
A8: A systems biology perspective of regulatory and metabolic adaptation of Staphylococcus aureus to infection-related conditions
Previous work showed that S. aureus responds rapidly and effectively to infection-relevant stimuli by adaptations of its gene expression and reorganization of protein complexes and metabolic pathways. For a comprehensive understanding of S. aureus physiology and virulence we will now elucidate adaptation of functional interaction networks with a focus on host challenges by a combination of experimental, bioinformatics and systems biology approaches. This reference model transcends the picture of static protein complexes by integrating the full dynamics of the regulatory and metabolic networks in S. aureus and will expand existing S. aureus pathogen models.
(German version)
A8: Eine systembiologische Analyse der metabolischen und regulatorischen Anpassung von Staphylococcus aureus auf infektionsbezogene Bedingungen
Wir zeigten bereits, dass S. aureus schnell und effektiv auf infektions-relevante Stimuli unter Anpassung seiner Genexpression, Stoffwechselleistungen und Proteinkomplexe reagiert. Für ein umfassenderes Verständnis der Physiologie und Virulenz von S. aureus werden wir nun die Anpassung funktioneller Interaktionsnetzwerke an wirtspezifische Faktoren durch eine Kombination von experimentellen, bioinformatischen und systembiologischen Ansätzen studieren. Dieses Referenzmodell geht über die Beschreibung der Proteinkomplexe hinaus und wird die Dynamik regulatorischer und metabolischer Netzwerke in S. aureus verbinden, um damit existierende S. aureus Modelle systembiologisch zu erweitern.
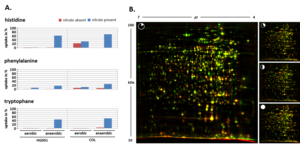
A more advanced understanding of regulation and metabolism in bacteria has to look beyond individual proteins, interactions and complexes to consider functional and regulatory networks by using a systems biological perspective. Recent examples of such systems biological studies include Mycoplasma pneumoniae and Bacillus subtilis. A metabolic S. aureus model has been established from our own and public transcriptome, proteome, and metabolome data to illustrate adaptation processes, including reorganization in central carbohydrate metabolism. The model shows that metabolic parameters such as redox state determine adaptive potential of S. aureus for the switch between commensal and pathogenic life style; secondary metabolism such as teichoic acid metabolism (collaboration with B5) was shown to be involved in strain evolution. Now we will elucidate regulatory and functional interaction networks in S. aureus in terms of system changes and adaption with a focus on host challenges combining experimental, bioinformatics and systems biology approaches. Physiological and gene expression experiments involving wild type strains and corresponding mutants lacking global regulators under different growth conditions will be used to validate this reference model (collaboration with A3 and B1). To reveal specific adaptation mechanisms involved in S. aureus virulence, omics data from complex growth scenarios such as cell culture and animal infection models (collaboration with C6, Z3) as well as from in vitro and in vivo biofilms (collaboration with A3, C6) will be included. This S. aureus model transcends the idea of static protein complexes and significantly expands existing pathogen models by an integrated systems biology perspective including host challenges and pathogen response and will be tested by targeted experiments (collaboration with A3, B1, C6).
Report and state of understanding
In the 1st funding period (A8 exists since 2010) we established and compared metabolic networks of different S. aureus strains (COL, N315, and Newman) [11] with a strong focus on differences in protein complexes [1], signalling switches and individual sequence mutations in regulatory elements (e.g. two component systems [22]). Protein complexes are rapidly changing in response to growth conditions and external stimuli and metabolic pathways display complex organization and regulation in S. aureus involving different levels and modules. Thus, a systems biology perspective is necessary for a more advanced understanding [2]. As evidenced from work on Mycoplasma [3] and B. subtilis [4] progressing the knowledge on individual protein complexes and metabolic pathways to a comprehensive systems description of all involved switches and network changes is a promising approach to get deeper insights into physiology and virulence. In the new period we want to combine the existing metabolic network models with regulatory networks and integrate them into our systems view, building on simple physiological adaptation scenarios with a new focus on host-pathogen interactions. In the following, we do not only report on the established data sets but show that these provide an excellent basis for the intended new project.
i. Omics data sets regarding metabolic adaptation
In close cooperation with other projects of the CRC-TRR34 we collected a large body of experimental data on the adaptive responses of S. aureus to different stress conditions. This includes transcriptomics, proteomics (including protein complexes [5]), and metabolomics data obtained under various infection-related growth conditions (e.g. starvation, exposure to oxidative stress, nitric oxide, heat, and antibiotics). In detail we analyzed the reorganization in central carbohydrate metabolism during glucose starvation [6] and the anaerobic gene expression of S. aureus [7, 8, 9]. A meta-study on anaerobiosis [8] revealed that S. aureus responds in different anaerobic experimental setups (anaerobic growth in the presence and absence of nitrate as terminal electron acceptor; anaerobic growth in the presence of supplemental pyruvate and uracil; anaerobic growth of an nreABC deficient mutant in the presence of nitrate) with a general anaerobiosis response. Among others, this response is characterized by an induction of several fermentation enzymes (PflB, Ldh1, SACOL0135, and SACOL0660) and the response regulator SrrA. Interestingly, especially genes with a high codon adaptation index highly overlap with anaerobically induced genes. With Rex the master regulator of the anaerobic gene expression in S. aureus was identified [9] (see section iii).
Additionally, we employed quantitative proteomics approaches considering subcellular localizations of more than 80% proteins expressed in growing and non-growing cells [5].
ii. Established metabolic networks
The already mentioned transcriptomics, proteomics, and metabolomics data were integrated by bioinformatics approaches to model adaptation processes [7, 10]. Comparing proteomics and metabolomics data we updated genome and proteome information and established strain-specific metabolic models for S. aureus COL, N315, and Newman including specific differences in metabolism and enzymes [11]. For instance, in glucose limitation experiments we found lower activity for enzymes in upper glycolysis and pentose phosphate pathway and stronger activity for some in lower glycolysis in exponentially growing S. aureus COL. In transition phase, aspartate kinase was expressed to meet amino acid requirements and in later phases there was high expression of glyceraldehyde-3-phosphate dehydrogenase and lysine synthetase. Furthermore, we provided a time-resolved picture of more than 500 proteins and 94 metabolites during the transition from exponential growth to glucose starvation. Under glucose excess, cells exhibited higher levels of proteins involved in glycolysis and protein synthesis, whereas entry into the stationary phase triggered an increase of enzymes of TCC and gluconeogenesis. These alterations in levels of metabolic enzymes were paralleled by more pronounced changes in the concentrations of associated metabolites, in particular, intermediates of the glycolysis and several amino acids [6].
iii. Systems biology and data integration
To plan and understand systems biology experiments in S. aureus and several other model organisms the GoSynthetic database (http://gosyn.bioapps.biozentrum.uni-wuerzburg.de) integrates data of one million processes, proteins, COGs and GOs [10]. The tool was applied to compare and map experimental data onto our genome-scale metabolic networks. The Aureolib database (http://www.aureolib.de) stores and compares thousands of time-dependent protein synthesis profiles of S. aureus COL under various infection-related growth conditions. This enabled us to define stimuli-specific marker proteins and to differentiate between growth rate related and stress specific gene expression [12].
From a systems biological perspective we concluded that intermediary metabolism mainly determines the adaptive potential of S. aureus for the switch between commensal and pathogenic life style [9, 13] while secondary metabolism was shown to be involved in strain evolution and resistance to antibiotics [5, 14, 15]. Importantly, there are central regulatory elements and their interaction partners orchestrating these metabolic adaptation processes. One of these is the redox-sensitive master regulator Rex. Its amino acid sequence and binding sequence (TTGTGAA–W4–TTCACAA) are highly conserved in all S. aureus strains. The binding activity of Rex is enhanced by NAD+ while NADH, which competes with NAD+ for Rex binding, decreases the activity of Rex. The impact of Rex on global protein synthesis and on the activity of fermentation pathways under aerobic and anaerobic conditions was analyzed by using a rex deficient strain [9].
iv. Adaptation of S. aureus to environmental challenges
In combined experimental and computational approaches we assessed not only effects on S. aureus induced by nutrient limitation but also effects of key regulators such as the quorum sensing network [16], sigB [17] and two component systems [22]. Combinations of different regulator mutations (agr, sae, sae /agr, sigB, sigB/sae) were tested in silico in a model and compared regarding system changes and correlation to experimental gene expression data. The model and simulations allow us to study quorum sensing mechanisms and biofilm formation in S. aureus including behaviour of MRSA strains and mutants [16]. The in vitro and in silico evidence highlights the role of sae and agr in fine-tuning biofilm repression and/or S. aureus dissemination. SigB was revealed to be a regulator but not a virulence determinant for the SigB-dependent transcription of virulence genes such as clfA, aur, and hla [17].
Complementary to these physiological studies we established a workflow and obtained time-resolved quantitative proteome profiling of Staphylococcus aureus HG001 internalised by human airway epithelial cells [13]. Moreover, we analyzed the response of S. aureus to the antibiotic mupirocin [20]. This compound selectively inhibits the bacterial isoleucyl-tRNA synthetase (IleRS) leading to the accumulation of uncharged isoleucyl-tRNA and the synthesis of (p)ppGpp. The alarmone (p)ppGpp induces the stringent response, an important global transcriptional and translational control mechanism that allows bacteria to adapt to nutritional deprivation. This includes global regulators such as CodY and SigB in shaping the response of S. aureus to mupirocin. Of particular interest was the induced transcription of genes encoding virulence-associated regulators (i.e., arlRS, saeRS, sarA, sarR, sarS, and sigB), as well as genes directly involved in the virulence of S. aureus (i.e., fnbA, epiE, epiG, and seb) [20]. Furthermore, we were also able to show that targeting primary metabolism may be more promising with less chances to adapt for S. aureus, xenobiotics such as lead compound IQ-143 and IQ-238 support this notion IQ-143 was bacteriostatic, and at higher concentrations bactericidal. Our analyses suggested that the mode of action was a direct interference in nucleotide and energy metabolism (apparent by metabolic modelling and gene expression analysis) with low toxicity in human cells [14]. An important source of resistance is that S. aureus clones exchange mobile genetic elements. This includes S. aureus pathogenicity islands (SaPIs) with high frequency via helper phages. Interestingly, horizontal gene transfer (HGT) has not been restricted to different S. aureus clones but SaPIs can be exchanged with other species and genera including Staphylococcus epidermidis and Listeria monocytogenes. This phenomenon was analyzed in close cooperation with the Peschel group (B5) for the S. aureus ST395 lineage, refractory to HGT with typical S. aureus. At the root of S. aureus ST395 separation is secondary metabolism of wall teichoic acid (WTA): ST395 produces an unusual WTA resembling that of its HGT partner species. Excitingly we could show that distantly related bacterial species and genera undergo efficient HGT with S. aureus upon ectopic expression of S. aureus WTA. Combined with genomic analyses, these results indicate that a 'glycocode' of WTA structures and WTA-binding helper phages permits HGT even across long phylogenetic distances thereby shaping the evolution of Gram-positive pathogens [15].
In conclusion, the established methods in data analysis and integration provided us important insights into the physiology and virulence of S. aureus. From the metabolic network modeled for different S. aureus strains general pathways and specific differences have been concluded. This together offers a great chance for the next funding period to extend our metabolic network models with regulatory information to reflect the natural situation where the metabolic state and transcriptional regulation are tightly connected, but even more so in response to host interaction and in pathogenic states. Profiting from the accumulated data of the CRC-TRR34 and our models and experiments the stage is set to achieve an integrated systems biology model of S.aureus in different states including infection settings.
References:
1. Krüger B, Liang C, Prell F, Fieselmann A, Moya A, Schuster S, Völker U, Dandekar T. 2012 Metabolic Adaptation and Protein Complexes in Prokaryotes. Metabolites 2: 940-958
2. Kim HD, Shay T, O'Shea EK, Regev A. 2009. Transcriptional regulatory circuits: predicting numbers from alphabets. Science 325:429-432
3. van Noort V, Seebacher J, Bader S, Mohammed S, Vonkova I, Betts MJ, Kühner S, Kumar R, Maier T, O'Flaherty M, Rybin V, Schmeisky A, Yus E, Stülke J, Serrano L, Russell RB, Heck AJ, Bork P, Gavin AC. 2012. Cross-talk between phosphorylation and lysine acetylation in a genome-reduced bacterium. Mol Syst Biol. 8:571
4. Nicolas P, Mäder U, Dervyn E, Rochat T, Leduc A, Pigeonneau N, Bidnenko E, Marchadier E, Hoebeke M, Aymerich S, Becher D, Bisicchia P, Botella E, Delumeau O, Doherty G, Denham EL, Fogg MJ, Fromion V, Goelzer A, Hansen A, Härtig E, Harwood CR, Homuth G, Jarmer H, Jules M, Klipp E, Le Chat L, Lecointe F, Lewis P, Liebermeister W, March A, Mars RA, Nannapaneni P, Noone D, Pohl S, Rinn B, Rügheimer F, Sappa PK, Samson F, Schaffer M, Schwikowski B, Steil L, Stülke J, Wiegert T, Devine KM, Wilkinson AJ, van Dijl JM, Hecker M, Völker U, Bessières P, Noirot P. 2012. Condition-dependent transcriptome reveals high-level regulatory architecture in Bacillus subtilis. Science 335:1103-1106
5. Becher D, Hempel K, Wolff S, Zühlke D, Pané-Farré J, Otto A, Fuchs S, Albrecht D, Bernhardt J, Engelmann S, Völker U, van Dijl J M, Hecker M. 2009. A proteomic view of an important human pathogen--towards the quantification of the entire Staphylococcus aureus proteome. PLoS ONE 4:e8176
6. Liebeke M, Dorries K, Zuhlke D, Bernhardt J, Fuchs S, Pane-Farre J, Engelmann S, Völker U, Bode R, Dandekar T, Lindequist U, Hecker M, Lalk M. 2011. A metabolomics and proteomics study of the adaptation of Staphylococcus aureus to glucose starvation. Mol Biosyst 7:1241-1253
7. Fuchs S, Pane-Farre J, Kohler C, Hecker M, Engelmann S. 2007. Anaerobic gene expression in Staphylococcus aureus. J Bacteriol 189:4275-4289
8. Fuchs S, Mehlan H, Kusch H, Teumer A, Zühlke D, Berth M, Wolf C, Dandekar T, Hecker M, Engelmann S, Bernhardt J. 2010. Protecs, a comprehensive and powerful storage and analysis system for OMICS data, applied for profiling the anaerobiosis response of Staphylococcus aureus COL. Proteomics. 16:2982-3000
9. Pagels M, Fuchs S, Pane-Farre J, Kohler C, Menschner L, Hecker M, McNamarra PJ, Bauer MC, von Wachenfeldt C, Liebeke M, Lalk M, Sander G, von Eiff C, Proctor RA, Engelmann S. 2010. Redox sensing by a Rex-family repressor is involved in the regulation of anaerobic gene expression in Staphylococcus aureus. Mol Microbiol. 76:1142-61
10. Liang C, Krüger B, Dandekar T. 2013. GoSynthetic database tool to analyse natural and engineered molecular processes. Database (Oxford) 2013:bat043
11. Liang C, Liebeke M, Schwarz R, Zuhlke D, Fuchs S, Menschner L, Engelmann S, Wolz C, Jaglitz S, Bernhardt J, Hecker M, Lalk M, Dandekar T. 2011. Staphylococcus aureus physiological growth limitations: insights from flux calculations built on proteomics and external metabolite data. Proteomics 11:1915-1935
12. Fuchs S, Zuhlke D, Pane-Farre J, Kusch H, Wolf C, Reiss S, Binh le TN, Albrecht D, Riedel K, Hecker M, Engelmann S. 2013. Aureolib - A Proteome Signature Library: Towards an Understanding of Staphylococcus aureus Pathophysiology. PLoS One 8:e70669
13. Schmidt F, Scharf S S, Hildebrandt P, Burian M S, Bernhardt J, Dhople V M, Kalinka J, Gutjahr M, Hammer E, Völker U. 2010. Time-resolved quantitative proteome profiling of host-pathogen interactions: the response of Staphylococcus aureus RN1HG to internalisation by human airway epithelial cells. Proteomics 10:2801-2811
14. Cecil A, Rikanović C, Ohlsen K, Liang C, Bernhardt J, Oelschlaeger TA, Gulder T, Bringmann G, Holzgrabe U, Unger M, Dandekar T. 2011. Modeling antibiotic and cytotoxic effects of the dimeric isoquinoline IQ-143 on metabolism and its regulation in Staphylococcus aureus, Staphylococcus epidermidis and human cells. Genome Biol. 12:R24
15. Winstel V, Liang C, Sanchez-Carballo P, Steglich M, Munar M, Penadés JR, Nübel U, Holst O, Dandekar T, Peschel A, Xia G. 2013. Wall teichoic acid structure governs horizontal gene transfer between major bacterial pathogens. Nature Communications 4:2345
16. Audretsch C, Lopez D, Srivastava M, Wolz C, Dandekar T. 2013. A semi-quantitative model of Quorum-Sensing in Staphylococcus aureus, approved by microarray meta-analyses and tested by mutation studies. Mol. BioSyst. 9:2665-80
17. Depke M, Burian M, Schäfer T, Bröker BM, Ohlsen K, Völker U. 2012. The alternative sigma factor B modulates virulence gene expression in a murine Staphylococcus aureus infection model but does not influence kidney gene expression pattern of the host. Int. J. of Medical Microbiology 302:33-39
18. Schoenfelder SM, Marincola G, Geiger T, Goerke C, Wolz C, Ziebuhr W. Methionine biosynthesis in Staphylococcus aureus is tightly controlled by a hierarchical network involving an initiator tRNA-specific T-box riboswitch. PLoS Pathog. 2013 Sep;9(9):e1003606
19. Somerville GA, Proctor RA. 2009. At the crossroads of bacterial metabolism and virulence factor synthesis in Staphylococci. Microbiol Mol Biol Rev. 73:233-48
20. Reiss S, Pane-Farre J, Fuchs S, Francois P, Liebeke M, Schrenzel J, Lindequist U, Lalk M, Wolz C, Hecker M, Engelmann S. 2012. Global analysis of the Staphylococcus aureus response to mupirocin. Antimicrobial agents and chemotherapy 56:787-804
21. Pragman AA, Yarwood JM, Tripp TJ, Schlievert PM. 2004. Characterization of virulence factor regulation by SrrAB, a two-component system in Staphylococcus aureus. J. Bacteriol. 186:2430-2438.
22. Krueger B, Friedrich T, Förster F, Bernhardt J, Gross R, Dandekar T. 2012. Different evolutionary modifications as a guide to rewire two-component systems. Bioinform Biol. Insights 6:97-128.