Molecular insights into spider silk
12/06/2018Spider silk belongs to the toughest fibres in nature and has astounding properties. Scientists from the University of Würzburg discovered new molecular details of self-assembly of a spider silk fibre protein.
They are lightweight, almost invisible, highly extensible and strong, and of course biodegradable: the threads spiders use to build their webs. In fact, spider silk belongs to the toughest fibres in nature. Based on its low weight it even supersedes high-tech threads like Kevlar or Carbon. Its unique combination of strength and extensibility renders it in particular attractive for industry. Whether in aviation industry, textile industry, or medicine – potential applications of this magnificent material are manifold.
Since long time material scientists continue to try reproducing the fibre in the laboratory, but with limited success. Today, it is possible to manufacture artificial spider silk of similar properties as the prototype, but the molecular-level structural details responsible for material properties await to be disclosed. Now, scientists from the Julius-Maximilians-Universität Würzburg (JMU) delivered new insights. Dr Hannes Neuweiler, lecturer at the Institute of Biotechnology and Biophysics at the JMU, is in charge of this project. His results are published in the scientific journal Nature Communications.
A molecular clamp connects protein building blocks
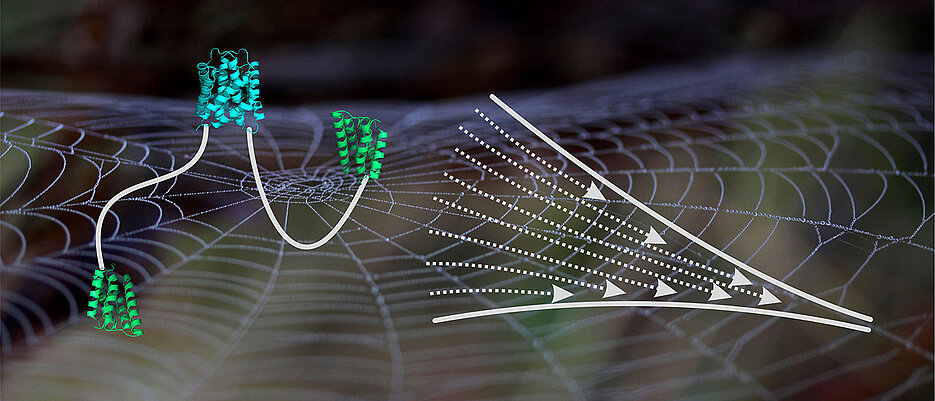
“The silk fibres consist of protein building blocks, so-called spidroins, which are assembled by spiders within their spinning gland”, explains Neuweiler. The terminal ends of building blocks take special roles in this process. The two ends of a spidroin are terminated by an N- and a C-terminal domain.
The domains at both ends connect protein building blocks. In the present study, Neuweiler and colleagues took a close look at the C-terminal domain. The C-terminal domain connects two spidroins through formation of an intertwined structure that resembles a molecular clamp. Neuweiler describes the central result of the study: “We observed that the clamp self-assembles in two discrete steps. While the first step comprises association of two chain ends, the second step involves the folding of labile helices in the periphery of the domain”.
This two-step process of self-assembly was previously unknown and may contribute to extensibility of spider silk. It is known that stretching of spider silk is associated with unfolding of helix. Previous work, however, traced extensibility back to the unfolding of helices in the central segment of spidroins. “We propose that the C-terminal domain might also act as module that contributes to extensibility” explains Neuweiler.
Assisting material science
In their study Neuweiler and co-workers investigated protein building blocks of the nursery web spider Euprosthenops australis. They used genetic engineering to exchange individual moieties of building blocks and modified the protein chemically using fluorescent dyes. Finally, the interaction of light with soluble proteins disclosed that the domain assembles in two discrete steps.
Neuweiler describes the result as “a contribution to our molecular-level understanding of structure, assembly and mechanical properties of spider silk”. It may aid material scientists to reproduce natural spider silk in the laboratory. Currently, modified and synthetic spidroins are being used for this purpose. “Should the C-terminal domain contribute to flexibility of the thread, material scientists may modulate mechanical properties of the fibre through modulation of the C-terminal domain”, Neuweiler says.
Two-step self-assembly of a spider silk molecular clamp. Charlotte Rat, Julia C. Heiby, Jessica P. Bunz & Hannes Neuweiler. Nature Communications. https://doi.org/10.1038/s41467-018-07227-5
Contact
Dr Hannes Neuweiler, Institute of Biotechnology and Biophysics, T: +49 931 31-83872, hannes.neuweiler@uni-wuerzburg.de