Virtual Platelet
We establish an interactive virtual platelet environment where research of different fields is integrated into a virtual agent-based platelet and megakaryocyte (MK) model that allows studying the spatial-temporal interactions that drive protein network signaling and cell decisions.
Our aim is to offer a space that allows user interaction with and manipulation of the environment. In particular, the user can explore and investigate the cell, and develop, plan, and test specific research questions.
For example, we introduce agent-based models for studying specifically the role of the cytoskeleton in platelet formation and signaling, exemplified by RhoA, RhoB, Cdc42 modulators of the central platelet signaling cascade.
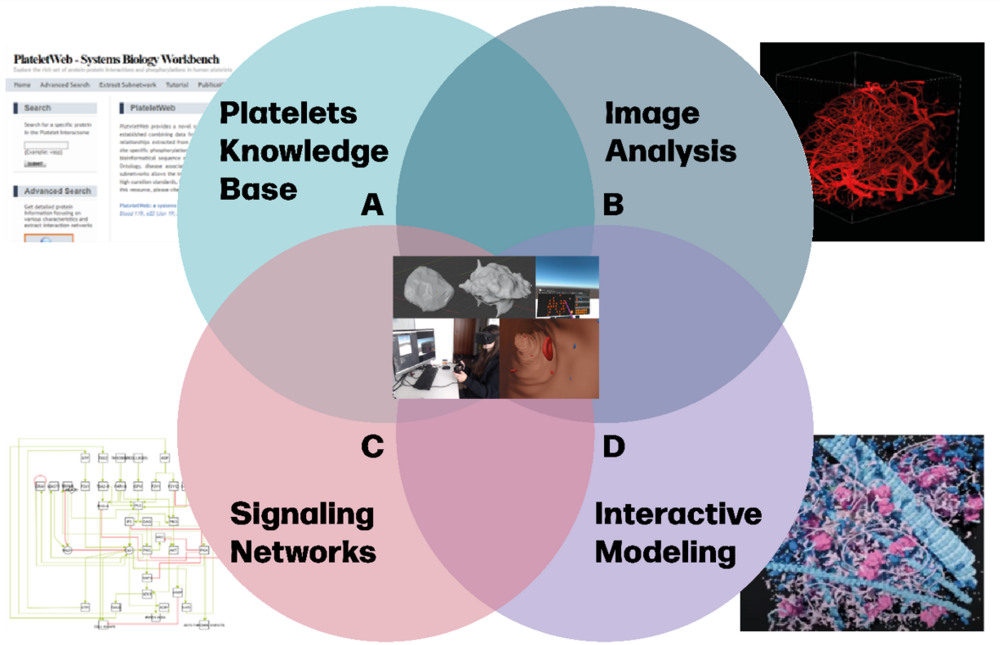
We will use different model scales resulting in an agent-based model based on an agent-based model within an interactive environment:
- Protein-protein interaction network signaling within platelet and megakaryocytes (MKs) driving platelet and MK behavior
- Cell-cell interactions of platelets and MKs on the tissue level, such as the platelet formation in the bone marrow or thrombus formation in brain vessels
- Within the virtual reality 3D world, the user dives into the platelet microcosmos and can interact with the simulation such as manipulating objects and simulation parameters and can retrieve information from the knowledge-base
- The VR simulations are based on scientific expertise, e.g. data warehouse on images of cells and tissues, as well as proteins, Boolean, and dynamic protein network models. Additionally, our experience in big data analysis (omics and image analysis) offers information and services on e.g., disease-related key proteins signaling, or feature detection in high-end microscopy images of platelets, MKs, bone marrow, or vessels
In sum, biology, imaging, and computer science (imaging, agent-based models, interactive simulation, virtual reality) are combined to establish an interactive virtual platelet environment that delivers new insights into the spatiotemporal models of platelets, MKs, and their cellular interactions.