P 6: M. Steinke
Characterization of host cell responses after Bordetella pertussis infection using 2D and 3D in vitro airway test systems
State of the art
Currently, mainly acellular vaccines are being used to prevent whooping cough caused by Bordetella pertussis. However, despite good vaccination coverage the disease incidence, particularly in infants, is strongly increasing in many countries in the past few years. Thus, the improvement of the current vaccines is mandatory requiring basic research on Bordetella pathogenicity mechanisms.
A major research problem with the etiological agents causing pertussis is the fact that B. pertussis and the close relative B. parapertussis are obligate human pathogens. Due to the use of at least partially inappropriate animal models (mainly mice) and cell culture model systems which do not resemble the natural situation, i.e. the ciliated and mucus-producing epithelium of the human respiratory tract, the real relevance of known virulence factors of B. pertussis for infection and disease still remains largely speculative. For example, the reasons for the major disease syndrome, the typical cough paroxysms, are not known, and still a debate is going on whether a so far not identified “cough toxin” remains to be discovered. Finally, very little is known so far about the host cell responses on infection which are involved in tissue pathology and clinical disease.
Previous Work
The Steinke group successfully generated a 3D in vitro test system of the human airway mucosa with high in vitro – in vivo – correlation (Fig A). Besides fully differentiated primary airway epithelial cells, the tissue model consists of fibroblasts that maintain the surrounding connective tissue and play an important role in basement membrane formation [1, 2]. Preliminary infection studies with both B. pertussis supernatants and bacteria led to cytoplasmic vacuoles, damaged mitochondria, cellular extrusions, and completely destroyed epithelial cells, which are associated with impaired epithelial barrier integrity (Fig. B).
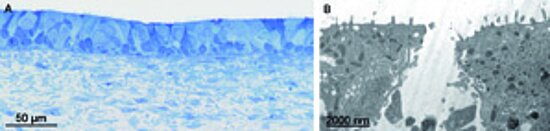

Figure: (A) Light micrograph of the intact airway mucosa test system (methylene blue staining), (B) destroyed airway epithelium after infection with B. pertussis supernatant (transmission electron microscopy).
The Gross group has a long-standing research experience with B. pertussis and closely related human and animal pathogens belonging to the genus Bordetella [1, 3-5]. Major research interests in the past were mainly focused on pathogenicity mechanisms of the bacteria, including the regulation of virulence gene expression in B. pertussis, invasion and survival of the bacteria in different cell types and the comparative characterization of virulence factors and genomes of B. pertussis and its close relatives.
Work Plan
B. pertussis specifically adheres to the ciliated epithelial cells of the airway mucosa. The production of virulence factors including several toxins causes dramatic changes in the epithelium such as extrusion of the ciliated cells from the tissue and massive mucus production. So far, very little is known about the signal transduction pathways involved and about the host cell response after infection in general. Signal transduction pathways are of particular importance since several toxins such as the pertussis toxin and the adenylate cyclase toxin are known to directly interfere with such pathways, but their contributions to clinical disease are not understood so far.
We hypothesize that the main cell types of the airway epithelium, namely goblet, ciliated and basal cells, respond differently to these virulence factors. Thus, the Steinke and Gross groups will apply well-characterized B. pertussis mutants lacking different virulence factors and cocktails of purified B. pertussis virulence factors to the airway mucosa in vitro test systems. The morphological impact of bacterial toxins such as the tracheal cytotoxin on the tissue models will be investigated by light and transmission electron microscopy (in cooperation with the Imaging Core Facility of the Biocenter).
The major scientific focus of our project will, however, be the host cell response upon infection. Using laser microdissection we will selectively collect goblet/ciliated/basal cells and fibroblasts from infected and non-infected tissues and perform transcriptome analyses. The analysis of the transcriptional response of the different cell types will show whether cell-type specific genetic programs are triggered which lead to previously described consequences of intoxication by B. pertussis such as necrosis of the ciliated cells and mucus hypersecretion by goblet cells.
In addition, we will investigate whether biological changes in living cells after B. pertussis infection can be monitored with Raman spectroscopy non-invasively. The 3D tissue model of the airway mucosa will be further developed by adding human microvascular endothelial cells on the basal side of the scaffold, which would simulate the endothelial barrier. In collaboration with the Schneider-Schaulies group focusing on measles virus transmission, we will extend the airway tissue models with infected and non-infected dendritic cells, which are considered of key importance in late viral transmission to epithelial cells.
References
- Steinke et al. (2014) An engineered 3D human airway mucosa model based on an SIS scaffold. Biomat 35:7355-7362. PubMed
- Rossi et al. (2014) Humane 3D-in-vitro-Testsysteme für die präklinische Forschung. Biospektrum DOI: 10.1007/s12268-014-0457-7. Biospektrum
- Bibova et al. (2015) Transcriptional profiling of Bordetella pertussis during infection reveals requirement of Hfq for type III secretion system functionality. RNA Biol 12(2):175-85. PubMed
- Yevsa et al. (2013) Development and characterization of attenuated metabolic mutants of Bordetella bronchiseptica for applications in vaccinology. Environ. Microbiol. 15:64-76. PubMed
- Gross et al. (2008) The missing link: Bordetella petrii is endowed with both the metabolic versatility of environmental bacteria and virulence traits of pathogenic Bordetellae. BMC Genomics 9:449. PubMed