P 1: A. Beilhack / M. Sauer
Host-pathogen interactions revealed by 3D high resolution microscopy
State of the art
Optical imaging technologies have revolutionized our perspective of the natural dynamics of biological and pathological processes. High-resolution microscopy of three dimensional environments has become powerful tools to dissect host-pathogen interactions. Advances in multi-photon laser-scanning microscopy, super-resolution microscopy and light-sheet fluorescence microscopy as well as improved fluorescent probes and genetically encoded flurorescent reporter genes offer a multifaceted toolbox to explore complex cellular and molecular processes in live tissues. Light-sheet microscopy, confocal and multi-photon laser-scanning microscopy methods have been optimized for observations of three dimensional environments with penetration depths of more than 500 micrometers.
Within the GRK 2157 we aim to improve and employ cutting-edge microscopy techniques to identify and exploit biological processes, decipher signalling pathways, and validate new drug targets. We will combine super-resolution microscopy (dSTORM/PALM) [1, 2] with light-sheet microscopy as an ideal tool to investigate cellular responses over time with high spatial resolution. We will develop set-ups to study host-pathogen interactions in 3D tissue models under conditions closely approximating those of a natural environment. The GRK consortium provides an ideal environment to foster interdisciplinary collaboration and highly innovative research to quantitatively elucidate host-pathogen interactions at unprecedented molecular, cellular, microanatomical and functional detail in a unique setting that mimics the onset and progression of infectious diseases in a human tissue environment.
Previous Work
The Beilhack group has been investigating immune cell interactions after allogeneic hematopoietic cell transplantation (HCT). Allogeneic HCT has proven as a successful treatment modality for patients with malignant diseases, immune deficiencies or severe autoimmune diseases. Yet, to date allogeneic HCT is facing three major challenges: First, adequate immune reconstitution and control of infections, second, overreaching immune responses that can result in life-threatening graft-versus-host disease and, third, impaired immune effector functions that are not sufficient to control the underlying disease, e.g. eliminating remaining leukemic cells. Thus, our research focuses on the interactions of the immune system with pathogens, cancer and the tissue environment [3-6]. To address these complex challenges of a dysfunctional immune system caught between defective tolerance and compromised immunity we employ optical imaging methods, i.e. optical techniques that provide single cell resolution in the context of intact tissues. We established dynamic multi-photon laser-scanning microscopy (MPM) that reveals complex cellular processes in real-time within tissues. Furthermore, we developed a novel multicolour light-sheet fluorescence microscopy (LSFM) approach for deciphering immune and infection processes in large tissue specimens on a single-cell level in 3 dimensions [7].
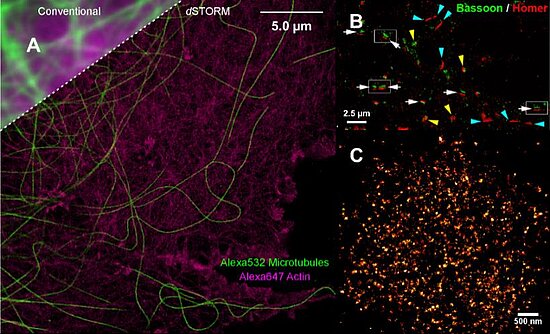
The Sauer group focusses its research on the development of new improved methods for the study of single biomolecules and their interactions with so far unmatched spatial and temporal resolution. Besides new methods for the study of conformational dynamics of biomolecules and protein folding, direct stochastic optical reconstruction microscopy (dSTORM) has been developed that enables super-resolution fluorescence imaging with standard organic fluorophores with a spatial resolution of ≤ 20 nm in the imaging plane in fixed and in living cells (Figure) [8].
Work Plan
We aim to employ MPM and LSFM in combination with dual-color PALM/dSTORM to 3D tissue models to study microbial infections by human pathogens. This will be achieved by an automated microscope performing imaging at virtually molecular resolution of a series of antibodies and nanobodies as well as other available specific probes. However, optical light-scattering and limited penetration depth still pose certain restrictions to this method. To date, we have overcome these by employing tissue clearing procedures. Yet, if tissue clearing is necessary, fixation of the tissues has limited real-time life cell imaging deep within tissues. Therefore, we want to optimize our microscopy approaches elucidating new NIR-dyes and improved clearing media as well as labeling protocols.
We will develop refined localization microscopy methods capable of providing quantitative information about molecular distributions and densities with so far unmatched spatial resolution. For example, the sub-cellular localization of bacterial proteins secreted by Chlamydia (Rudel), Salmonella (Vogel) and Campylobacter (Sharma) and their interacting host proteins will be determined. To generate maps of interacting proteins with high spatial resolution, we will use quantitative dSTORM/PALM [9]. The development of quantitative localization microscopy tools will be accompanied by correlative electron microscopy and array tomography experiments.
References
- Heilemann et al. (2008). Subdiffraction-resolution fluorescence imaging with conventional fluorescent probes. Angew Chem Int Ed 47, 6172–6176. PubMed
- van de Linde et al. (2011). Direct stochastic optical reconstruction microscopy with standard fluorescent probes. Nat Protocols 6, 991–1009. PubMed
- Rieber et al. (2015). Pathogenic fungi regulate T-cell immunity by inducing neutrophilic yeloid-derived suppressor cells through a Dectin-1/CARD9 and ROS-mediated mechanism. Cell Host Microbe 8;17(4):507-14. PubMed
- Donat et al. (2012). Surface display of Gaussia princeps luciferase allows sensitive fungal pathogen detection during cutaneous aspergillosis. Virulence 3, 50-61. PubMed
- Chopra et al. (2013). Tumor necrosis factor receptor 2-dependent homeostasis of regulatory T cells as player in TNF-induced experimental metastasis. Carcinogenesis 34, 1296-1303. PubMed
- Vaeth et al. (2015). Selective NFAT targeting in T cells ameliorates GvHD while maintaining anti-tumor activity. Proc Natl Acad Sci U S A 112, 1125-1130. PubMed
- Brede et al. (2012) Mapping immune processes in intact tissues at cellular resolution. J Clin Invest, 122: 4439-4446. PubMed
- Wombacher et al. (2010). Live-cell super-resolution imaging with trimethoprim conjugates. Nat Methods 7, 717–719. PubMed
- Ehmann et al. (2014). Quantitative Super-Resolution Imaging of Bruchpilot Distinguishes Active Zone States. Nat. Communications 5, 4650-4661. PubMed