Focus of Research
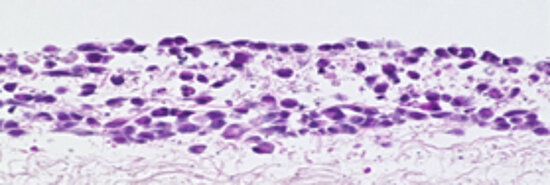
We aim to elucidate the molecular and mechanistic basis for interactions between host and microbes in natural infections with the long-term goal to develop new anti-infective strategies. Our approach fosters the study of infections in 3D human tissue models reflecting the natural infection site in humans. The main entrance routes for human pathogens are the skin as well as the respiratory, gastrointestinal, and urogenital tract. Engineered 3D human tissues of these entry routes will be utilized for infection experiments with selected human-specific microbes (Chlamydia, Neisseria, Campylobacter, Bordetella, Salmonella, Trypanosoma and measles virus).
We plan to establish vascularized tissue models to address bacterial dissemination, such as seen in gonococcal infection, whereas models for secondary barriers including human endothelia, 3D human blood brain barrier or dynamic DC/T cell interactions will be developed with groups working with microbes causing meningitis or encephalitis (meningococci, trypanosomes and measles virus). Natural tissues consists of more than one cell type, and certain cell types may be selectively engaged during traversal of microbes through infection barriers such as the epithelial layer. Moreover, individual cells within natural tissues may behave differently from cells within monolayers, and may even be metabolically reprogrammed in response to pathogen encounter. These predispositions not only require the investigation of microbes during interactions with complex tissue, but also the exploration of the response of individual host cells to the infection.
The challenges of performing molecular analysis in such complex infection models will be met by applying the latest next-generation and high-throughput technology for two general read-outs: 1) bioimaging (super-resolution fluorescence imaging, dSTORM, PALM, in vivo fluorescence imaging, BLI etc.), and Raman Spectroscopy will be used to obtain overall optical and physical information of cell behaviour; 2) single- and multi-cell next-generation RNA- and DNA-sequencing combined with bioinformatics analysis will provide information on gene expression changes during the course of infection.
Besides the discovery of the basic principles of infections and validation or disproval of data generated with “classical” model systems, the identification of infection-induced signalling pathways, changes in cellular trafficking, cell-cell transmission or other infection related phenotypes will likely yield new targets for the development of therapeutic strategies to protect from or combat bacterial, parasite and viral infections.