Scientific Environment
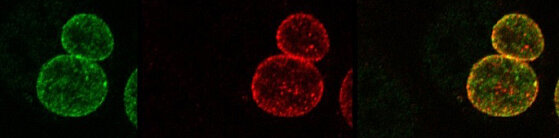
The work performed at the Chair of Tissue Engineering and Regenerative Medicine has offered a new perspective on infection models for obligate human pathogens. For example, human tissue can be built by colonizing an acellularized matrix of porcine gut with a monolayer of differentiated, polarized human cells. These models can be built from different cell types and can even be vascularized. Infection biologists have collaborated with Heike Walles and her team to evaluate the feasibility of using engineered tissue as a model for infections with their respective pathogens (Jannasch et al., 2015).
For example, infections with Bordetella pertussis in a human trachea model (Steinke et al., 2014), with the food-borne pathogen Campylobacter jejuni in a small intestine model (C. Sharma & H. Walles, unpublished), and the eukaryotic pathogen Trypanosoma b. gambiense in a human full thickness skin model (M. Engstler & H. Walles, unpublished) have been successfully performed. These initial experiments have already clearly demonstrated that infection of engineered tissue can provide fascinating insight into conditions mimicking the natural tissue.
A spectacular experiment was the injection of Trypanosomes by the tsetse fly into a 3D human skin model. In further infection experiments the 3D vascularized skin model will be used to study if and how the trypomastigotes may eventually access the blood stream simulating natural infection mechanisms. Thus, we have successfully generated tissue models including the human airway mucosa, intestinal mucosa and skin, which represent the major barrier organs and closely reflect the cellular complexity found in vivo (Schanz et al., 2010). In addition, to organize reliable material sources for engineered tissues based on primary cells, a cell bank and network of collaborating surgeons in the University hospital of Würzburg and other hospitals has been established since 2009 under ethical approval.
These pilot experiments, however, also uncovered the challenges that have to be met during the development of these infection models. These include the integration of immune cells, functional vascularization and partly allogeneic cell components. A major limitation of the currently established models is the lack of an intact immune system in the engineered tissue. We will therefore undertake a major effort to test how cells of the innate or adaptive immune system can complement the models under development (see project S. Schneider-Schaulies).
Another challenge to be approached is the adaptation of analytical methods to complex 3D tissue cultures. To this purpose, several already established techniques can be used. Imaging of the multilayer tissue in real time, at high resolution and to very high depth can be achieved by dynamic multi-photon laser-scanning and multiparameter light-sheet fluorescence microscopy (A. Beilhack), super-resolution microscopy (M. Sauer) and combinations thereof, as well as established 3D-high speed fluorescence microscopy with water immersion objectives (M. Engstler). Regulation of gene expression in the pathogen and the host in response to infection can be monitored by conventional or/and dual RNA-Seq. Since the response of either the pathogen or the host may differ depending on the cell type that is infected, RNA-Seq may have to be performed on single or a small numberof cells isolated from mixed populations (J. Vogel). Changes in gene expression will not only be monitored, but functional assays using RNA interference-mediated gene silencing (in host cells) or gene knockout (in host and pathogens) will be applied, as well. Lentivirus-mediated delivery of short hairpin RNAs allows for silencing gene expression also in primary cells (T. Rudel, S. Schneider-Schaulies) whereas CRISPR/Cas9 knockout technology can be applied in immortalized cells (J. Vogel). A very important key discipline for the success of image and sequencing analyses is bioinformatics-based data mining combined with systems biological modeling (T. Dandekar). Modeling of 3D cell culture and infection barriers such as the blood-brain barrier is currently being performed.
These projects are embedded in an international and widely visible research environment within the University of Würzburg.
The GRK 2157 collaborates with the:
- Graduate School of Life Sciences (GSLS)
- CRC/TRR 124 "FungiNet - Pathogenic fungi and their human host:
- Networks of interaction"