Genome studies: more is not always better
07/14/2021The characteristics of plants of the same species can have different genetic causes depending on their origin. This is shown by a recent study at the University of Würzburg.
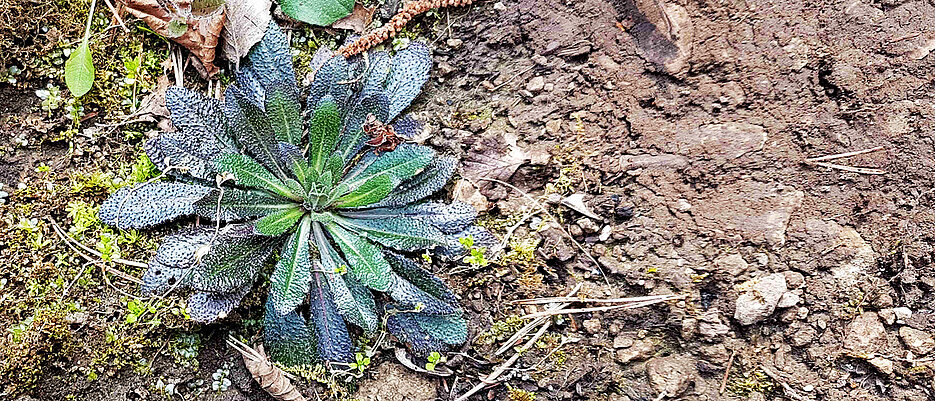

What the fruit fly is to zoologists, the thale cress is to botanists. The widespread herb with the botanical name Arabidopsis thaliana serves them as a model organism from which knowledge can be gained for other plants. It is therefore extremely well researched - also genetically. For example, it is now known that the genetic material of Arabidopsis thaliana (its genome) comprises around 125 million base pairs. It's like having a Lego manual in front of you that is 125 million letters long and contains everything you need to know to build an Arabidopsis plant.
Similar to humans, different Arabidopsis specimens are generally not genetically identical. If you were to compare the construction manual of all plants of this species, you would encounter differences in about 10 million places, experts estimate. "We have now taken a closer look at three million of these variable sites in the genome," explains Arthur Korte, junior professor of Evolutionary Genomics at the University of Würzburg. "And we did so in nearly 900 Arabidopsis plants from very different locations across Europe, from southern Spain to central Sweden."
For botanists, the variations in the genome are very interesting. because they are responsible for differences between individual Arabidopsis plants. Some plants can, for example, cope better with drought, while others are more resistant to frost. "To some extent, these are traits that we would like to introduce into our crop plants," Korte explains. "But to do that, we first need to know which genetic differences are related to which traits in the plant."
Too much heterogeneity is harmful
Classically, scientists use a method known by the abbreviation "GWAS" (genome-wide association study) for this purpose. They examine the genomes of thousands of plants and look for changes in the genetic blueprint that are particularly frequently associated with certain traits, such as better drought resistance. The more specimens one compares in this way, the more such links between genotype (the individual genetic blueprint) and phenotype (the properties of the plant in question) should stand out.
"But we were able to show in our study that this is not necessarily the case," Korte emphasizes. "Instead, it's sometimes better to limit yourself to fewer specimens from a defined local area." The reason: plant populations that grow in locations with very different conditions often differ significantly in their genomes. This heterogeneity might lead to the scenario that a trait such as drought resistance has very different genetic origins in different locations. "Therefore, if a GWAS includes many plants with very high genetic heterogeneity, it may miss important associations between genotype and phenotype," Korte says.
In their study, the scientists were indeed able to demonstrate this effect. On the one hand, they conducted a GWAS of all nearly 900 plants. In addition, they also examined only subpopulations - for example, those Arabidopsis specimens that had been collected on the southern Iberian Peninsula. "In doing so, we found genetic correlations that were not seen in the overall population because they had become too diluted there," says Korte. "These results show that valuable new insights can be gained from smaller, more genetically homogeneous samples." That applies not only to plants, by the way, but just as much to GWAS in humans.
Local adaptations are often based on changes in gene networks
The study also provides interesting insights into the evolution of new traits: genetic adaptations to local conditions (for example, to a particularly dry environment) are usually not based on the fact that a single "drought gene" has changed and thus become more effective. Instead, they often involve regulator genes, which in turn intervene entire networks of traits. "These regulators could then provide a better fine-tuning of already existing metabolic pathways," says Korte.
This finding is also relevant for breeding new varieties. In the past, it was often thought that one simply had to cross a certain gene into a breeding line in order to obtain the desired trait there. In the meantime, however, it has become increasingly clear that networks of many different factor have to be considered. "We are now learning better and better how to identify such networks," says Korte. “With this knowledge, it might be possible in the future to adapt today's crops to new challenges such as climate change. "
Publication
William Andres Lopez-Arboleda, Stephan Reinert, Magnus Nordborg, Arthur Korte, Global genetic heterogeneity in adaptive traits, Molecular Biology and Evolution, 2021; https://doi.org/10.1093/molbev/msab208
Contact
Arthur Korte, Juniorprofessur für evolutionäre Genomik, Center for Computational and Theoretical Biology (CCTB), University of Würzburg, T: +49 931/31-80361, arthur.korte@uni-wuerzburg.de