A damage foreseen is a danger avoided
11/11/2020New complex for damage detection in DNA identified. Our body can repair damage to our DNA that can lead to the development of cancer by means of repair complexes. But how does the repair machinery recognize the damage? Scientists from the University of Würzburg and the University of Kent have now identified a complex that plays an important role in damage recognition in nucleotide excision repair. Due to its key position, the complex represents a starting point for research on cancer drugs. The results were published in the renowned journal Nucleic Acids Research.
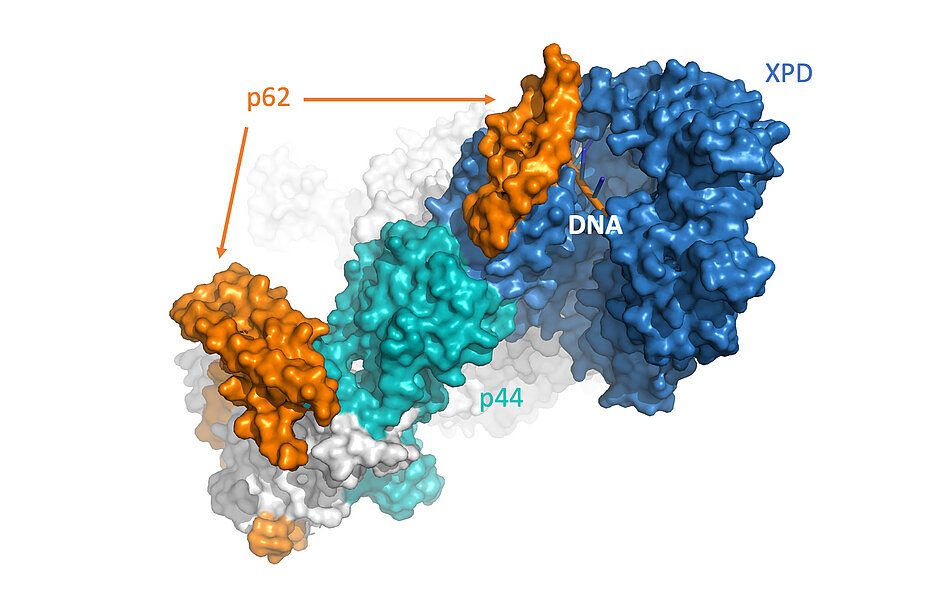
For a cell to survive, it is extremely important that it can repair DNA damage, such as those caused by UV radiation or chemicals, quickly and efficiently. The cell has various repair mechanisms at its disposal. A well-known mechanism is nucleotide excision repair (NER), which utilizes a large protein complex consisting of ten subunits. The interaction of three of those subunits namely XPD, p44 and p62 makes an essential contribution to the detection of damaged DNA, as was demonstrated by the research groups of Prof. Dr. Caroline Kisker from the Rudolf Virchow Center for Integrative and Translational Bioimaging at the University of Würzburg and Dr. Neil Kad from School of Biosciences at the University of Kent. XPD is already used in medicine as a biomarker to predict the individual effectiveness of specific drugs in patients. The current study helps the researchers to better understand how the repair complex works.
XPD specifically detects UV damage
One highly important subunit in the NER repair complexes is the enzyme XPD. XPD separates the two DNA strands from each other during repair and is involved in damage verification. Eventually, XPD provides the signal for the removal of the defective DNA. This task is regulated and supported by the subunits p62 and p44. "Until now, it has been assumed that p62 primarily has a stabilizing function in the complex. With our experiments, we are now showing that this subunit assumes an additional function and contributes to better binding of the enzyme to the DNA," explains Kisker. p62 apparently also interacts directly with the DNA and simultaneously enhances the activity of the XPD enzyme. "The effect of better binding can be observed especially in damaged DNA, but only if all three: XPD, p44 and p62 are present," says Kad. This enables the enzyme to detect detrimental changes in the DNA and enables effective repair.
NER as target structure for cancer drugs
Some cancer drugs aim to damage the DNA of cancer cells. If the repair mechanism is now disabled, the cells can no longer repair this damage and die. For this reason, researchers are looking for ways to block the NER specifically with inhibitors. A further advantage is that DNA damage in cancer cells that are no longer repaired by the NER system can activate the body's own immune response and thus contribute to fighting the tumor cells. Research into this complex and how it works will therefore continue to play an important role in the future.
Publication
Barnett JT, Kuper J, Koelmel W, Kisker C and Kad NM The TFIIH subunits p44/p62 act as a damage sensor during nucleotide excision repair Nucleic Acids Research (November 2020) doi: 10.1093/nar/gkaa973
Persons
Prof. Dr. Caroline Kisker is, among other things, head of the Department of Structural Biology and Dean of the Graduate School of Life Sciences at the University of Würzburg. Since May 2020, she is the speaker of the Rudolf Virchow Center for Integrative and Translational Bioimaging at the University of Würzburg. More information: https://www.uni-wuerzburg.de/en/rvz/lehrstuehle/lehrstuhl-fuer-strukturbiologie/
Dr. Neil Kad is the head of a research group at the University of Kent investigating DNA repair mechanisms.
Contacts
Prof. Dr. Caroline Kisker (Rudolf Virchow Center, University of Würzburg)
Tel.: +49 (0)931 31 80381, caroline.kisker@virchow.uni-wuerzburg.de
Dr. Neil Kad (University of Kent)
Tel.: +44 (0)1227 (8)16151, n.kad@kent.ac.uk
Dr. Judith Flurer (Press office, Rudolf Virchow Center)
Tel.: +49 (0)931 31 85822, judith.flurer@virchow.uni-wuerzburg.de