Prof. Dr. A. Westermann
Research
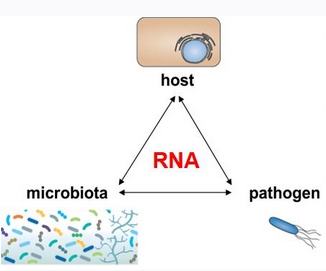
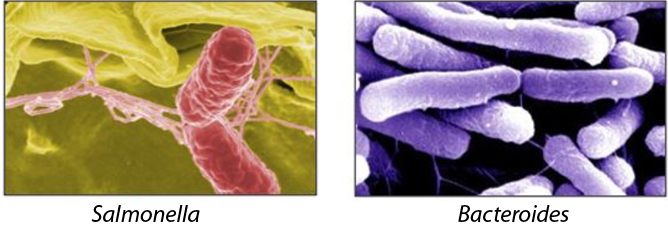
Our intestinal tract offers an attractive environment for both beneficial and pathogenic bacteria. The beneficial bacteria of our microbiota feast on undigested foods and provide numerous health benefits. Enteric pathogens see this environment as an entry point for infection. Both groups influence each other, creating a tripartite interaction with us, the host. Understanding the regulatory processes that decide on the outcome of these encounters represents an emerging research area to combat infectious diseases. While the field has focused on protein-mediated processes, our group investigates the role of RNA-centric mechanisms in controlling microbial interactions in the gut.
Ryan D, Bornet E, Prezza G, Varshini Alampalli S, Franco de Carvalho T, Felchle H, Ebbecke T, Hayward R, Deutschbauer AM, Barquist L, Westermann AJ (2024)
An expanded transcriptome atlas for Bacteroides thetaiotaomicron reveals a small RNA that modulates tetracycline sensitivity
Nature Microbiology 9(4):1130-1144
Prezza G, Liao C, Reichardt S, Beisel CL, Westermann AJ (2024)
CRISPR-based screening of small RNA modulators of bile susceptibility in Bacteroides thetaiotaomicron
PNAS 121(6):e2311323121
Westermann AJ*,+, Vogel J*,+ (2021)
Cross-species RNA-seq for deciphering host-microbe interactions
Nature Reviews Genetics 22(6):361-378
Ryan D, Jenniches L, Reichardt S, Barquist L, Westermann AJ (2020)
A high-resolution transcriptome map identifies small RNA regulation of metabolism in the gut microbe Bacteroides thetaiotaomicron
Nature Communications 11(1):3557
Westermann AJ, Förstner KU, Amman F, Barquist L, Chao Y, Schulte LN, Müller L, Reinhardt R, Stadler PF, Vogel J (2016)
Dual RNA-seq unveils noncoding RNA functions in Salmonella-host interplay
Nature 529(7587):496-501
Jun Prof. Alexander Westermann
![[Translate to Englisch:] [Translate to Englisch:]](/fileadmin/_processed_/1/5/csm_Westermann_Alexanderl_6570d77986.jpg)
Elise Bornet
Taís Franco de Carvalho
Hoda Kooshapour
Gohar Mädler
![[Translate to Englisch:] [Translate to Englisch:]](/fileadmin/_processed_/d/b/csm_Gohar_M_23426df8c5.jpg)
Sarah Reichardt
![[Translate to Englisch:] [Translate to Englisch:]](/fileadmin/_processed_/d/7/csm_Sarah_R_5e73f5764b.jpg)
Gianluca Prezza
![[Translate to Englisch:] [Translate to Englisch:]](/fileadmin/_processed_/4/3/csm_Gianluca_P_acb57472a7.jpg)
Dr. Daniel Ryan
![[Translate to Englisch:] [Translate to Englisch:]](/fileadmin/_processed_/6/9/csm_Daniel_R_d7b2823d7b.jpg)
Ann Sophie Rüttiger
![[Translate to Englisch:] [Translate to Englisch:]](/fileadmin/_processed_/0/c/csm_Ann-Sophie_R_6eb2cb23ad.jpg)