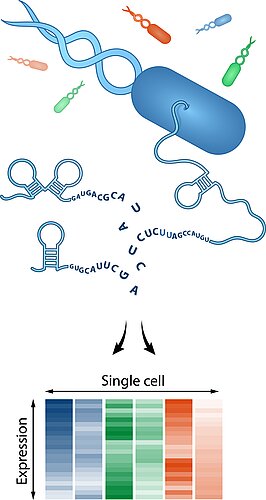
Single-cell RNA biology
The revolutionary ability of single-cell RNA-seq to obtain gene expression profiles of individual cells rather than to look of average expression in populations has led to profound new discoveries in biology, including the identification of new cell types. Single-cell RNA-seq is also emerging as a powerful tool to determine cell-to-cell variability in bacterial pathogenesis but for technical reasons, this technique has been restricted to the analysis of eukaryotic transcriptomes. For example, we have applied it to infected macrophages to study how heterogeneous intracellular growth of Salmonella drives their polarization state.
The grand challenge for microbiology now is to also establish single-cell RNA-seq for bacteria, species that lack a poly-A tail for easy cDNA priming as in eukaryotes, and contain even less RNA to begin with. Transcriptomes from single bacteria will help to better understand phenotypic resistance to antibiotic treatment, stochastic persister formation under harsh conditions, and phenotypic diversification for bet hedging, to name a just a few application. lt would also allow for single-cell Dual RNA-seq in which pathogen and host are analyzed simultaneously.
Beyond simple single-cell transcriptomics, it's our ambition to adapt other powerful RNA-seq techniques tobe able to study posttranscriptional control in single bacteria. For example, single-cell UV CLIP-seq would teil us about cell-to-cell variation of RNA-protein interactions. To this end, we are building expertise in designing microfluidics devices, for example, to perform RNA co-immunoprecipitation with proteins in minute reaction volumes. The long-term goal is high-throughput RNA biochemistry in single bacterial cells.