New RNA-bindings proteins
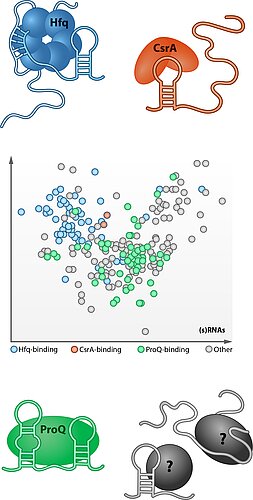
Noncoding RNA molecules typically interact with RNA-binding proteins (RBPs) in order to function in a cellular environment. Decades of work in eukaryotes have established RBPs as a pillar of posttranscriptional gene control, with numbers ranging in the hundreds to thousands in vertebrate and yeast cells. lronically, we know little about the number and mechanisms of RBPs in those organisms that seem the easiest to analyse: Bacteria. Although bacteria amply regulate their genes on the RNA level, our knowledge of global RBPs remains limited to the sRNA chaperone Hfq and the translational repressor CsrA. However, increasing evidence including the serendipitous discovery of CRISPR-Cas systems suggests that many RBPs await discovery in microbes.
We have discovered the "osmoregulatory" protein ProQ as a new abundant RBP with many cellular targets in Salmonella and E. coli. Our results for ProQ promise new molecular mechanisms of bacterial gene control at mRNA 3' ends and a novel regulatory network wherein ProQ recognizes RNA structure rather than primary sequence. We are also addressing how other members of this RBP family use the FinO domain for either highly specific or global RNA recognition in the same cellular environment.
Continuing with RBP discovery, we are now exploring the microbiome for new RBPs, using both multispecies Grad-seq and complementary approach akin to eukaryotic RNA-protein interactome screens.